Complex I (NADH dehydrogenase)
Introduction:
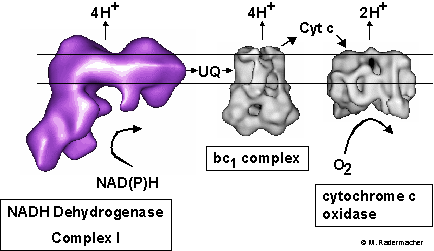 Model of the respiratory chain. In eukaryotes complex I oxidizes NADH present in the mitochondrial matrix. In this process protons are pumped into the outer membrane space of the mitochondrion and two electrons are transferred to ubiquinone (UQ). The reduced ubiquinol is then transferred to the bc1 complex (III) which again pumps protons to the outer membrane space and reduces cytochrome c (Cyt c). In cytochrome c oxidase (IV) cytochrome c is finally oxidized, again in connection with proton pumping. As a result of this process the matrix of the mitochondrion has a negative potential relative to the cytosol. This electroosmotic potential drives the ATP synthase (complex V) which phosphorylates ADP to give ATP, which is transported out of the mitochondrion via an ADP/ATP antiporter. (see e.g. Rasmusson, et al., 1998) )
Model of the respiratory chain. In eukaryotes complex I oxidizes NADH present in the mitochondrial matrix. In this process protons are pumped into the outer membrane space of the mitochondrion and two electrons are transferred to ubiquinone (UQ). The reduced ubiquinol is then transferred to the bc1 complex (III) which again pumps protons to the outer membrane space and reduces cytochrome c (Cyt c). In cytochrome c oxidase (IV) cytochrome c is finally oxidized, again in connection with proton pumping. As a result of this process the matrix of the mitochondrion has a negative potential relative to the cytosol. This electroosmotic potential drives the ATP synthase (complex V) which phosphorylates ADP to give ATP, which is transported out of the mitochondrion via an ADP/ATP antiporter. (see e.g. Rasmusson, et al., 1998) )
Structure and Function:
Our laboratory studies the structure and function of complex I by three-dimensional electron microscopy (see also Radermacher et al. Journal of Structural Biology, 154, 2006, 269-279). Complex I is a highly flexible structure and only an extensive classification of images into classes of identical particles made it possible to reconstruct a detailed structure. We used Random conical tilting for single particle 3D reconstruction.
From 35 images one can construct 3×1038 movie sequences. We selected 15 images that show the central membrane arm protuberance moving from front to back/back to front and constructed by hand a highly speculative movie of possible conformational changes of complex I: Watch the movie.
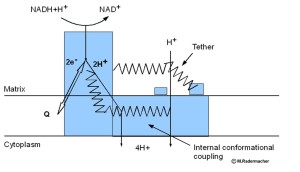
Our early model of a conformationally driven proton pumping mechanism of complex I. (s. also Meth. Enym. 2009, 456, 3-27)
In some 2D averages of Y. lipolytica complex I and in some intermediate 3D reconstruction a faint line can be observed, that could indicate a very thin connection between the Matrix arm domain (5) and the membrane arm protrusions, assumed to be located near proton pumping subunits. This tether may be one way to transfer conformational changes in the Matrix arm to the membrane arm. For a conformationally driven complex I two pathways for conformational coupling may exist. Either through the tether, or through more internal conformational changes. The recent X-ray structures of complex I from E. coli and from T. thermophilus (Efremov et al. Nature, 2010, 465, 441-445, Baradaran et al. Nature 2013, 494,443-448) showed an unusual long horizontal helix in the membrane arm, that may be involved in the proton pumping mechanism.
The overall structure of complex I is preserved throughout species.
While most of the early published structures of complex I showed essentially an L-shaped structure there seemed to be a contradiction in in the basic shape of the complex. Mostly published structures of bovine complex I and the E.coli complex I seemed to contradict the generally accepted shape of the enzyme. One of the major problems for determining the structure of the complex by electron microscopy is its high flexibility. This requires special attention and the use of techniques that can reliably separate these conformational variations. In addition to the structure of complex I from Y.lipolytica, we determined the structures of bovine complex I and complex I from A.aeolicus and were able to show that the basic domain structure of complex I is preserved across species ( T. Clason et al. J Struct Biol (2010) 169(1):81-88)
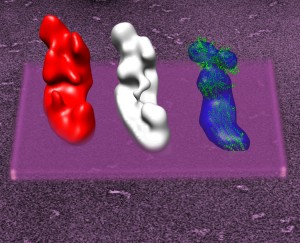
Structures of complex I. From left to right: Complex I from Y. lipolytica, bovine complex I, bacterial complec I (A. aeolicus).
Selected Literature:
M. Radermacher, Chapter 1: Visualizing Functional Flexibility by Three-Dimensional Electron Microscopy: Reconstructing Complex I of the Mitochondrial Respiratory Chain. In: Methods in Enzymology, Mitochondrial Function, Part A: Mitochondrial Electron Transport Complexes and Reactive Oxygen Species, Edited by: William S. Allison and Immo E. Scheffler, (2009) Vol. 456, 3-27
Angerer H, Radermacher M, Małkowska M, Steger M, Zwicker K, Heide H, Wittig I, BrandtU, Zickermann V. The LYR protein subunit NB4M/NDUFA6 of mitochondrial complex I anchors an acyl carrier protein and is essential for catalytic activity. Proc Natl Acad Sci U S A. 2014 Apr 8;111(14):5207-12.
Radermacher M, Ruiz T, Clason T, Benjamin S, Brandt U, Zickermann V.The three-dimensional structure of complex I from Yarrowia lipolytica: a highly dynamic enzyme. J Struct Biol. 2006 Jun;154(3):269-79.
T. Clason, V. Zickermann, T. Ruiz, U. Brandt, M. Radermacher, Direct Localization of the 51 kDa and 24 kDa Subunits of Mitochondrial Complex I by Three-dimensional Difference Imaging, Journal of Structural Biology, (2007), 159(3):433-442.
Brandt U. Energy converting NADH:quinone oxidoreductase (complex I). Annu Rev Biochem. 2006;75:69-92.
Abdrakhmanova A, Zickermann V, Bostina M, Radermacher M, Schagger H, Kerscher S, Brandt U.Subunit composition of mitochondrial complex I from the yeast Yarrowia lipolytica.Biochim Biophys Acta. 2004 Jul 23;1658(1-2):148-56.
Zickermann V, Bostina M, Hunte C, Ruiz T, Radermacher M, Brandt U. Functional implications from an unexpected position of the 49-kDa subunit of NADH:ubiquinone oxidoreductase. J Biol Chem. 2003 Aug 1;278(31):29072-8.
Peng G, Fritzsch G, Zickermann V, Schagger H, Mentele R, Lottspeich F, Bostina M, Radermacher M, Huber R, Stetter KO, Michel H. Isolation, characterization and electron microscopic single particle analysis of the NADH:ubiquinone oxidoreductase (complex I) from the hyperthermophilic eubacterium Aquifex aeolicus. Biochemistry. 2003 Mar 18;42(10):3032-9.
Djafarzadeh R, Kerscher S, Zwicker K, Radermacher M, Lindahl M, Schagger H, Brandt U. Biophysical and structural characterization of proton-translocating NADH-dehydrogenase (complex I) from the strictly aerobic yeast Yarrowia lipolytica. Biochim Biophys Acta. 2000 Jul 20;1459(1):230-8.
Rasmusson AG, Heiser V V, Zabaleta E, Brennicke A, Grohmann L.Physiological, biochemical and molecular aspects of mitochondrial complex I in plants Biochim Biophys Acta. 1998 May 6;1364(2):101-11.

